Implementation of Rhizomes
There are at least three different ways how to implement a rhizome on existing hard- and software platforms: as an object tree, relying on a programming language pointer arithmetic, or using mathematical pairing functions.
Implementation as object tree
The most obvious option is to re-interpret a rhizome as an object tree and use an object-oriented approach to implementation. This is probably the easiest way to quickly come to a working solution. This works for a relatively small number of relations, but not for big numbers.
Implementation based on pointer arithmetic
A more sophisticated option is to use a programming language with pointer arithmetic capabilities such as C/C++ or C#. This was indeed Erez Elul’ and Miriam Bedoni’ choice when implementing the first version of their Pile Engine. (I personally prefer to use the term rhizome for pile as I deem it more expressive.) There is also another version by Ralf Westphal (?), which I think is different from the first. Unfortunately, it is hard to find more information other than a few only partially functional links:
Although this is a highly viable option, it can become quite tricky to solve the implementation details, for example to parallelize the whole implementation.
Implementation based on pairing functions
There is however a third, highly interesting alternative, described in a short blog post by Ralf Barkow. The idea is to use a mathematical pairing function for our implementation. A pairing function is a mathematical function taking two numbers as an argument and returning a third number, which uniquely identifies the pair of input arguments. It is always possible to re-compute the pair of arguments from the output value. Two pairing functions are currently known to me.
Cantor pairing function:
pair(x, y) = z = 1/2 * (x^2 + 3x + 2xy + y + y^2)
unpair(z) = (x, y) = {
x = z - (q * (1 + q))/2,
y = (q * (3 + q))/2 - z },
with q = floor((-1 + sqrt(1 + 8z))/2)Szudzik pairing function:
pair(x, y) = z = {
x^2 + x + y, if x = max(x, y)
y^2 + x, otherwise
}
unpair(z) = (x, y) = {
x = z - floor(sqrt(z))^2, y = floor(sqrt(z)), if z - floor(sqrt(z))^2 < floor(sqrt(z))
x = floor(sqrt(z)), y = z - floor(sqrt(z))^2 - floor(sqrt(z)), otherwise
}Using one of the above pairing functions, we can now assign non-negative integer values to relators and relata. The relator rk is assigned the pairing value z, and the relata (ri, rj) the ordered pair of values (x, y). According to our definition, terminal relations are defined by “pointing to themselves”, in other words both relata being the same: rk <= (ri, rj) with ri = rj. Thus, it is strictly guaranteed that terminal relations are the only ones where x = y.
This figure shows an example for the Szudzik pairing function.
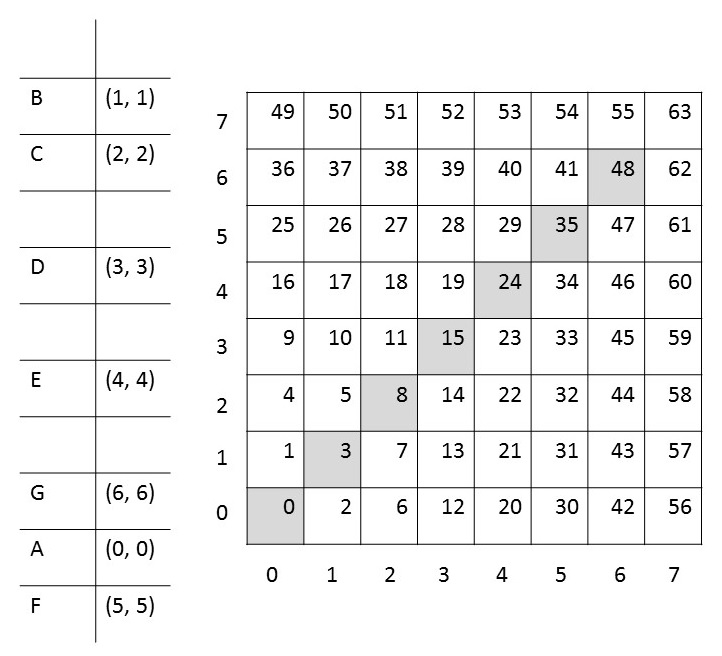
There are seven terminal relations (0, 0) to (6, 6) representing the characters A to G. The grid shows the z pairing values for each ordered pair (x, y) according to the Szudzik pairing function. Let us assume we wanted to encode the String ABC as two nested ordered pairs (A, (B, C)).
- Replace all characters with their terminal ordered pair of values:
((0, 0), ((1, 1), (2, 2))). - Repeat replacing all ordered pairs with their z pairing value until there is only a single z value left: 1)
(0, (3, 8)), 2)(0, 67), 3)4489.
The relator r4489 encodes the nested ordered pairs (A, (B, C)). Nesting and order do matter, the result would be different for ((A, B), C) or (A, (C, B)).
Of course the process works in a reverse way too.
- Repeatedly expand all
zpairing values by replacing them with an ordered pair of(x, y)values until you meet a pair wherex = y: 1)4489, 2)(0, 67), 3)((0, 0), (3, 8)), 4)((0, 0), ((1, 1), (2, 2))). - Replace all ordered pairs by looking up their data items in the dictionary:
(A, (B, C)).
We can now traverse the rhizome upwards and downwards (from relata to relator and vice versa) without the need to store any data. The grid shown above only describes an algorithm how to pair values, it is not stored anywhere as real data content. The only place so far where actual data is stored is for the mapping of atomic symbols to terminal relations. This is very attractive because the symbol/terminal-dictionary grows only linearly with an increasing number of terminals.
So how do we keep track of which non-terminal relations have already been established and which have not? We know for example how to compute the z-pairing value for the pair of symbols (B, C), but how can we know if such a pairing has been established or not? For this purpose, we need another data structure. Its only task is to keep track of what pairings have been established and which have not.
Edit 18th August 2015: I have found a probably even more efficient way of storing pairings than described below. See my blog entry Implementation of Rhizomes - Part 2.
The simplest approach is to use a two-dimensional bit array equivalent to the logical grid above. If a bit is set in the 2d-array, this indicates that a connection exists between its indices x and y. This will result in a matrix which is quite densely populated for low x and y values but sparsely populated for high x and y values. Unfortunately, the matrix grows in size at O(n2). Even if we only store bits this can quickly eat up all our resurces. At least for the sparsely populated matrix area we should try to find leaner solution.
As an alternative, I suggest using one more hashmap (or possibly list) structure. The hashmap contains each relatum appearing in a relation as a key, and two lists as its value. One list contains all associative relata for this (normative) relatum, the other contains all normative relata for this (associative) relatum. The list themselves are stored as packed bit strings. The best bit string packing algorithms do not even require the bit string to be unpacked to check whether a certain bit is set or not, only for setting or deleting a bit unpacking and repacking is executed. (Daniel Lemire’s blog contains some articles on bit packing in C++ and Java.) The list should be able to quickly return a list of relata this relatum is in a relation with. An algorithm I used personally is the CONCISE algorithm (Colantonio and Di Pietro [1]), which seems to even beat Word Aligned Hybrid (WAH) bitmap compression [4] on which for example the FastBit library [5], [6] is based. Colantonio wrote a highly efficient open source Java implementation of the CONCISE algorithm called Extendedset, which is available in two GitHub repositories [2], [3].
Some remarks
Some general remarks. First, using pairing functions only works if all terminals are assigned ordered pairs where ri = rj. In the grid, all terminals are stored along the grid’s diagonale. The reason is that a stop criterion is required when traversing the rhizome in a reverse way from relator to relata. Imagine that we would allow terminals to be stored also outside the diagonale. Then, for example, the character C could be stored as (0, 2). As both 0 and 2 appear as z-values (relators) in the grid, we cannot decide if the ordered pair (0, 2) already represents a terminal or must be unpaired even further to ((0, 0), (1, 0)).
Second, the z pairing values can become quite large. With a linear growth of the x and the y values, the z values grow quadratically. Yet, at the same time, the expressive power or the information content of z-values also grows quadratically. At the same time, the need for larger z-values continuously decreases, as certain complicated combinations of terminals rarely occurr.
Third, this approach requires to decide upon a predefined set of terminals. It is impossible to decide a posteriori to split a terminal into several sub-relations. Whereas this sounds like a severe restriction, I believe that it is not on many cases. In most real-world examples there exists a naturally restricted vocabulary or set of atomic symbols, which we silently accept.
- For example in digital computing, the atomic symbols are the binary digits
0and1. Or, taking a byte as an atomic symbol, the set of possible bytes contains only 256 different atoms. - In language, character sets such as ASCII or UTF-8 provide a large number of atomic symbols letter and number symbols which are expressive enough for the vast majority of use cases.
- Natural language uses a limited - albeit large - number of words. Applying rhizomes on natural language processing we could actually create a combined set the most common words enhanced with the atomic letter and number symbols to create less common words.
- The periodic table consists of a few dozens of chemical elements.
- The human genomes consists of only four fundamental molecules, the guanine (G), adenine (A), thymine (T) and cytosine (C).
Fourth, because rhizome trees do not store data items but only relations, multiple dictionaries can be applied on the same rhizome tree at the same time. In other words, the interpretation of any z-value or ordered pair (x, y) is entirely the responsibility of the user and the context. The situation is the same to processing binary data. The interpretation of a bit string like 00001010 is entirely left to the computational context, which not only defines whether this bit string is to be read from left to right or from right to left but also whether it designates a character, number, color, screen position, i/o address or something else. The minimum requirement for each context is a dictionary mapping atomic symbols to terminal relations and a second map or list keeping track of the actual pairings.
References
[1] Colantonio A., Di Pietro R. (2010): CONCISE: Compressed ‘n’ Composable Integer Set. Paper submitted to arXiv.org. No. 1004.0403.
[2] Extendedset: An open source Java implementation of the CONCISE algorithm authored by Alessandro Colantonio. Version 1.
[3] Extendedset: An open source Java implementation of the CONCISE algorithm authored by Alessandro Colantonio. Version 2.
[4] Wu K., Otoo E. J., Shoshani A. (2001): A Performance Comparison of bitmap indexes. Proceedings of the tenth international conference on information and knowledge management (CIKM ‘01). p. 559-561
